6 Module 6: Microbial Genome Sequencing
6.1 Overview
Weeks 11 and 12 focus on DNA isolation, quantification, sequencing preparation, and early genome analysis.
6.2 Purpose
- Evaluate DNA sequencing results and microbial genome assemblies.
- Follow a standard DNA isolation protocol.
- Apply bioinformatics tools to assemble microbial genomes and analyze predicted coding sequences.
6.3 Learning Outcomes
- List safety considerations for extracting DNA from microbial isolates.
- Explain the purpose of reagents used for DNA extraction.
- Discuss the significance of high-molecular-weight DNA for sequencing.
- Practice extracting DNA from liquid cultures.
- Practice quantifying DNA with a Qubit fluorometer.
- Describe the purpose, methods, and preliminary results of DNA extraction, quantification, and sequencing experiments.
- Collect and interpret DNA quantification and sequencing data.
- Revise the draft for the individual and group projects.
- Explain how the experiment contributes to the overall research goal.
6.4 Skills and Knowledge
6.4.1 Skills
- Follow lab safety and PPE procedures.
- Extract and quantify microbial DNA.
- Analyze microbial DNA sequence data.
6.4.2 Knowledge
- PPE requirements for DNA isolation and purification.
- DNA extraction and quantification methods.
- Oxford Nanopore Technologies sequencing workflows.
6.5 Task
Review the protocol before lab and work with your partner to isolate DNA from overnight cultures in duplicate, record all steps, and document the results.
6.6 Criteria for Success
Successful completion requires participation in DNA isolation and sequencing preparation, collection of analyzable data, and a complete ELN entry.
6.7 Background
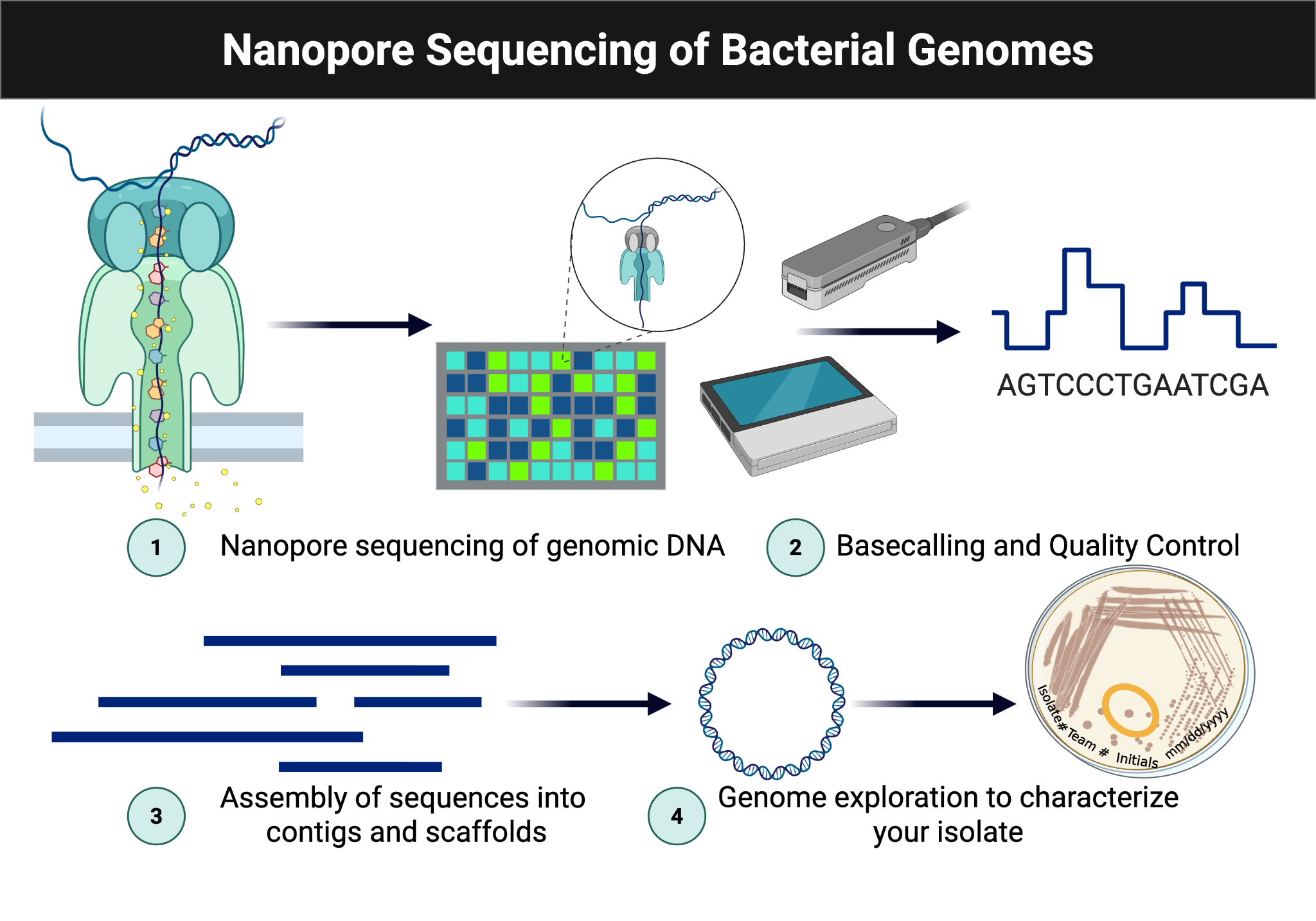
This module connects phenotype to genotype by sequencing the genome of the isolate. DNA is extracted using the NO-MISS workflow and analyzed with Oxford Nanopore sequencing and downstream genome analysis tools such as BV-BRC and SeqHub.
Figure Figure 6.1 summarizes the sequencing and downstream analysis pipeline from isolate DNA to annotated genome.
6.8 Procedures
6.8.1 Lab Safety
- Wear lab coat, goggles, and gloves.
- Clean benches and pipettors with 70% ethanol.
- Treat all culture-associated materials as biohazards.
- Decontaminate liquids, plates, and work areas at the end of the session.
6.8.2 Methods
6.8.2.1 What Was Prepared in Advance
- Overnight cultures were started in TSB.
- Lysozyme was prepared in TE buffer at 10 mg/mL.
6.8.2.2 Day of the Lab
- Pellet 1.0 mL of culture in a DNA LoBind tube.
- Resuspend in TE buffer with lysozyme.
- Add Proteinase K and RNase A.
- Add CTAB buffer and incubate at 56 C for 30 minutes.
- Combine lysate with Monarch gDNA Binding Buffer at a 2:1 ratio.
- Bind DNA to the spin column, wash twice, and elute with preheated Monarch gDNA Elution Buffer.
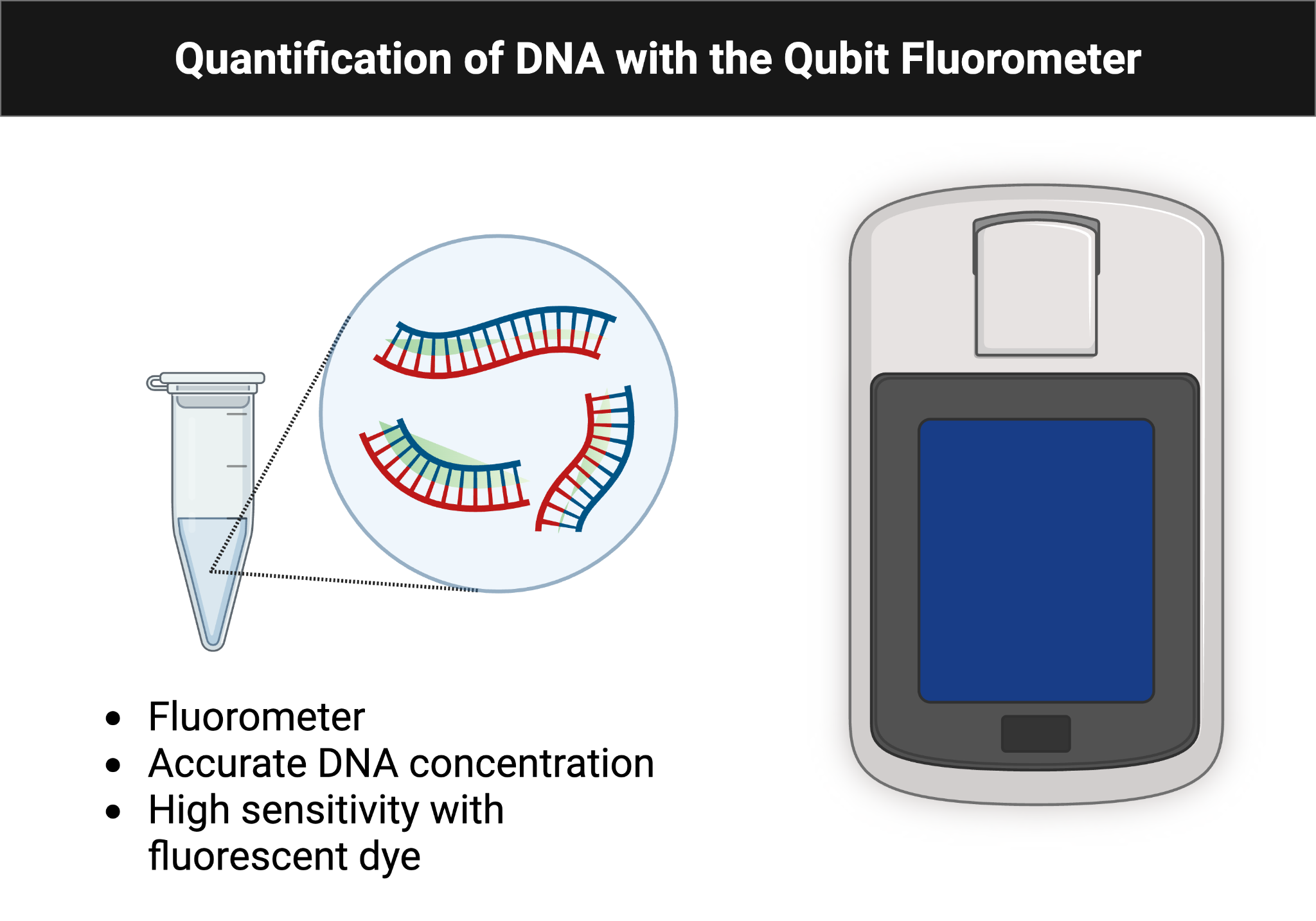
- Quantify 1 µL of DNA with the Qubit fluorometer.
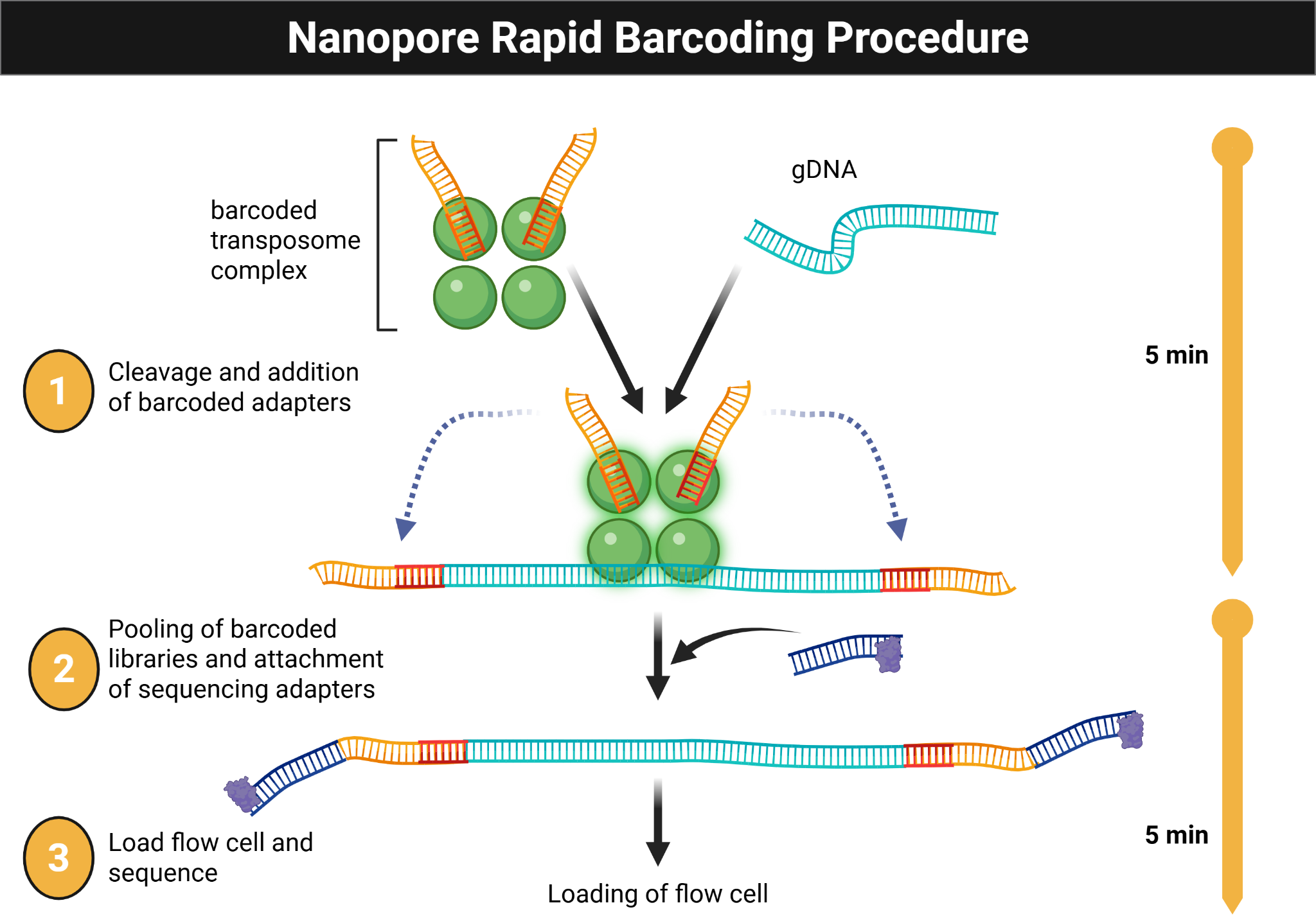
- Prepare sequencing-ready DNA according to the NO-MISS rapid barcoding workflow.
Figure Figure 6.2 illustrates the Qubit measurement setup used to quantify extracted DNA prior to library preparation.
Figure Figure 6.3 summarizes the rapid barcoding library-preparation workflow used before sequencing.
6.8.3 Data Analysis
- Create a BV-BRC account and access the shared workspace.
- Obtain the Nanopore and Illumina files for your isolate.
- Run the Comprehensive Genome Analysis workflow.
- Upload the assembled FASTA file to SeqHub for annotation and exploration.
6.8.4 Protocol Notes
Record any mistakes, deviations, or strain-specific observations.
6.9 Results
Add figures, screenshots, concentration values, and any sequencing outputs generated from your data.
6.10 Result Analysis
Discuss DNA yield, sample quality, and whether the sequencing-related results matched expectations. Include factors that may have affected DNA recovery and what you would change next time.
6.11 Discussion Questions
- How do microbial genome assembly and annotation tools support research?
- What makes nanopore sequencing different from other sequencing approaches?