3 Module 3: Microbial Growth Curves
3.1 Overview
Weeks 4 and 5 focus on growth-curve experiments, multichannel pipetting, and biofilm viability assays.
3.2 Purpose
- Evaluate growth rates of different Delftia species.
- Prepare liquid cultures from single colonies for 96-well growth assays.
- Analyze growth parameters from shaking cultures.
- Investigate differences in biofilm formation among isolates.
3.3 Learning Outcomes
- List safety considerations for propagating microbes.
- Explain the differences between static and shaking growth.
- Describe the features of biofilms.
- Discuss the significance of growth curves.
- Practice using a multichannel pipette.
- Practice diluting overnight cultures and seeding 96-well plates consistently.
- Describe the purpose, methods, and preliminary results of microbial propagation experiments.
- Collect and interpret growth and biofilm data.
- Create an annotated bibliography with at least five sources.
3.4 Skills and Knowledge
3.4.1 Skills
- Work safely with liquid cultures in 96-well plates.
- Document protocols and results clearly.
- Interpret growth-curve and biofilm data.
- Propagate bacteria in liquid culture.
3.4.2 Knowledge
- PPE requirements for microbial work.
- Propagation of microorganisms in liquid media.
- Seeding 96-well plates for growth and biofilm analyses.
3.5 Task
Review the procedures before lab and work with your partner to complete the growth and biofilm assays and document all observations.
3.6 Criteria for Success
Successful completion requires participation in both assay workflows, collection of analyzable data, and a complete ELN entry.
3.7 Background
This module compares growth and biofilm formation in Delftia acidovorans, Delftia tsuruhatensis, and student isolates while maintaining strong contamination control.
3.8 Procedures
3.8.1 Lab Safety
- Wear the required PPE.
- Clean the bench and pipettors with 70% ethanol.
- Treat all tips as biohazards.
- Dispose of plates and gloves in the correct waste streams.
3.8.2 Methods: Liquid Culture Setup
- Isolated colonies were transferred to liquid medium.
- Inoculated tubes were grown overnight at 30 C with shaking at 200 to 250 RPM.
3.8.3 Methods: Growth Assay Setup
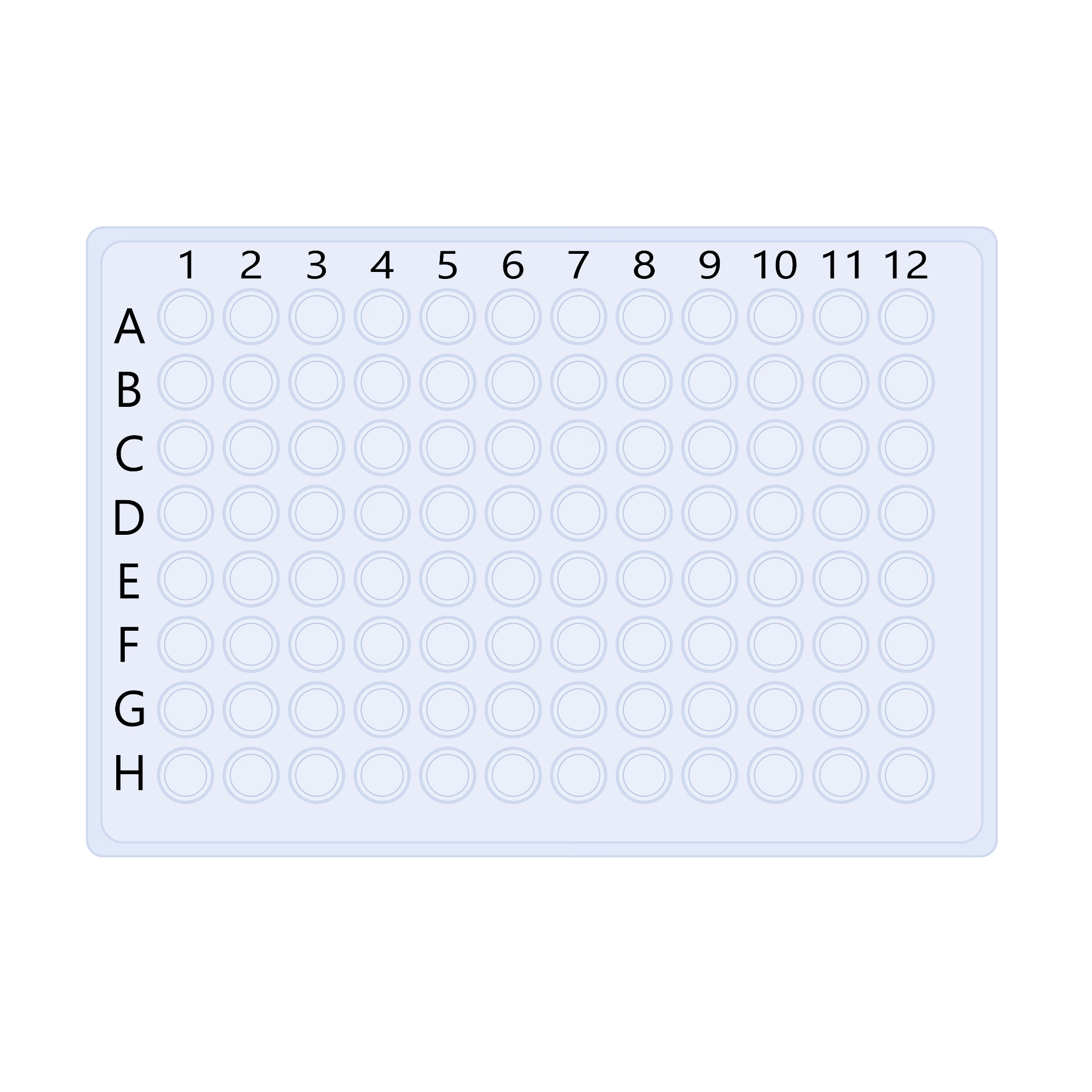
Figure Figure 3.1 provides a reusable 96-well template for mapping control wells and isolate conditions before loading the plate.
- Prepare 1:1000 dilutions of the overnight cultures.
- Use positive and negative controls.
- Pour diluted cultures into sterile reservoirs.
- Use a multichannel pipette to add 100 µL to the assigned 96-well plate columns.
- Seal the plate with a breathable seal.
- Load the plate into the plate reader for 48 hours at 30 C with shaking.
3.8.4 Methods: Biofilm Assay Setup
- Prepare fresh 1:1000 dilutions for the biofilm plate.
- Seed the assigned wells with 100 µL per well.
- Add the peg lid without contaminating the pegs.
- Incubate for 48 hours at 30 C.
3.8.5 Methods: Biofilm Viability
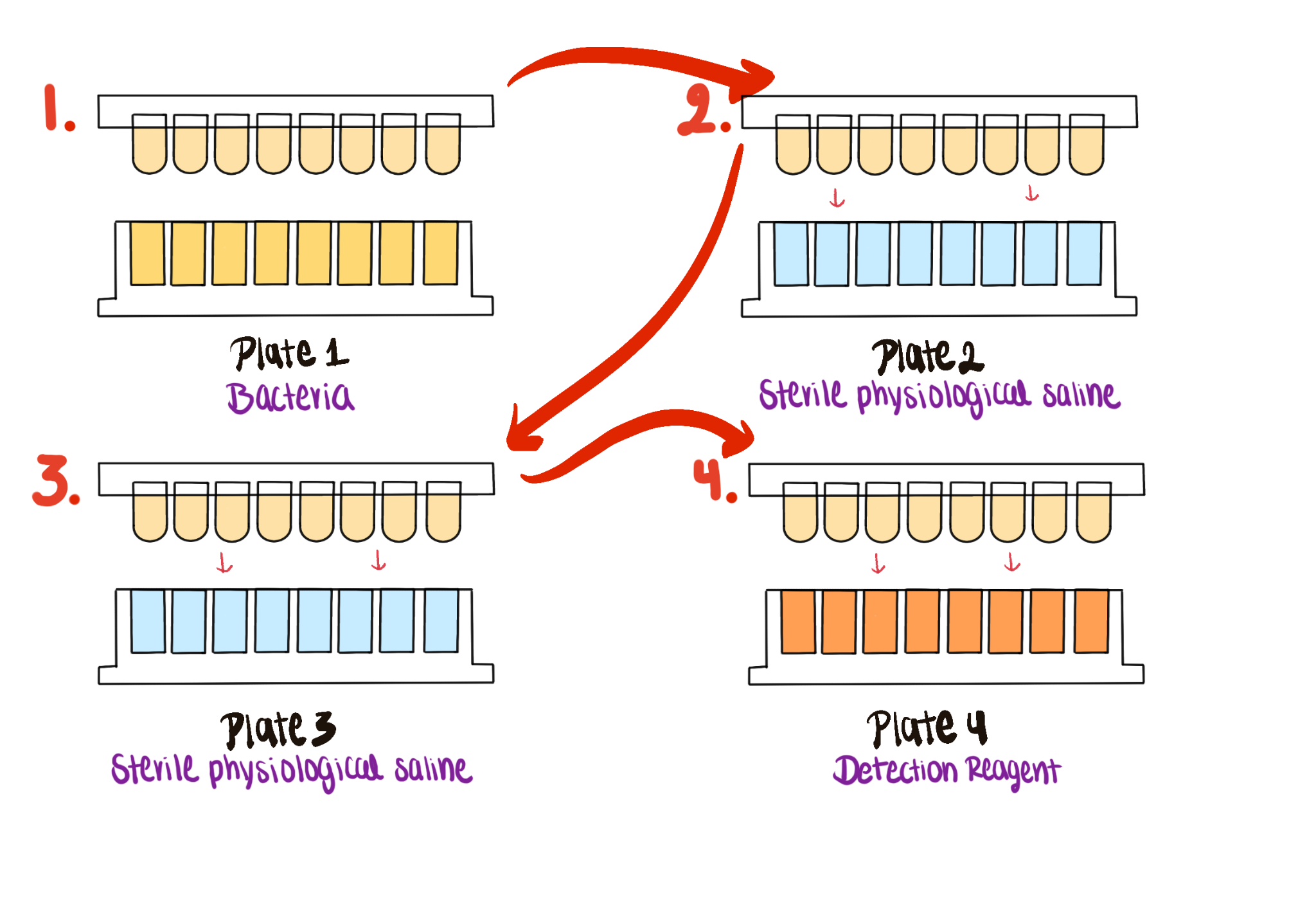
Figure Figure 3.2 outlines the wash-and-detection workflow used to measure biofilm viability after incubation.
- Wash the peg lid using PBS in separate plates.
- Prepare the detection reagent with WST solution, electron mediator reagent, and TSB.
- Transfer the peg lid to the reagent plate and incubate at 37 C.
- Measure absorbance at 450 nm.
3.8.6 Protocol Notes
Record any mistakes, deviations, or isolate-specific observations.
3.9 Results
3.9.1 Growth Curves
- Save and share the raw plate-reader data.
- Format the data for analysis.
- Upload the data file to the repository.
- Plot growth curves and compare isolates and wells.
3.9.2 Biofilm Viability
- Save and share the raw absorbance values.
- Subtract the average TSB-only background.
- Calculate average absorbance and standard deviation for each condition.
- Plot the background-corrected averages for comparison.
3.10 Result Analysis
Use your code and plots to explain whether the observed growth and biofilm behavior matched expectations.
3.11 Discussion Questions
- Why are biofilms difficult to eliminate, even when antibiotics are used?
- What growth-curve pattern would you expect for your isolate, and why?